Design and validation of PIP gene primer for quantitative PCR in Capsicum annuum using in silico and experimental approaches
DOI:
https://doi.org/10.23917/bioeksperimen.v12i1.16129Keywords:
C. annuum, Design Primer, PIP Gene, qPCRAbstract
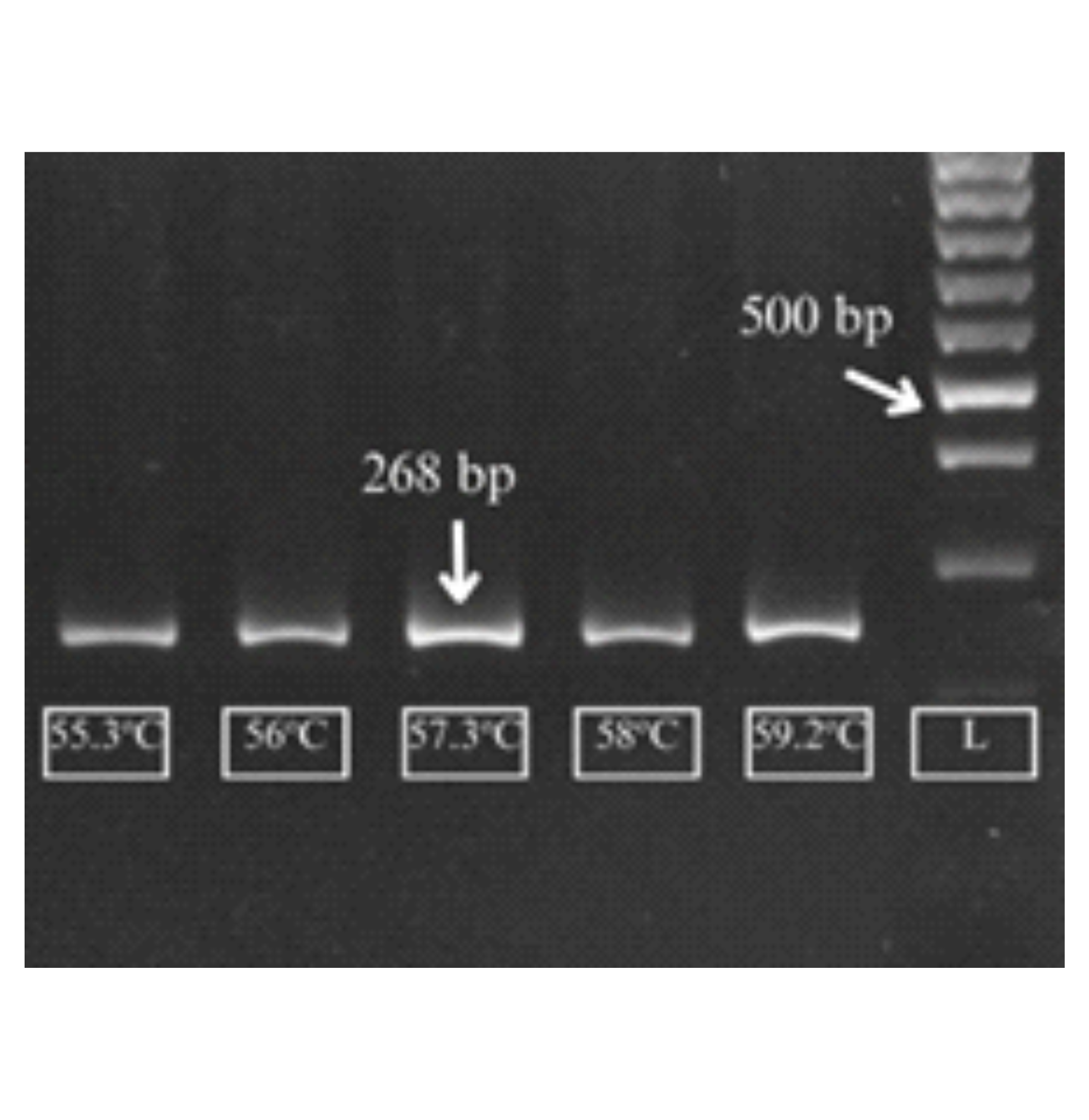
Capsicum annuum L. is an economically important horticultural crop whose productivity is strongly affected by drought stress. Plasma Membrane Intrinsic Protein (PIP), a member of the aquaporin gene family involved in water transport and osmotic regulation, represents a key target for drought stress studies, requiring highly specific and efficient primers for accurate gene expression analysis using quantitative PCR (qPCR). This study aimed to design and validate PIP gene primers for qPCR in C. annuum using integrated in silico and experimental approaches. Primer design was performed using NCBI Primer-BLAST based on the CaPIP reference sequence (XM_016711608.2), followed by in silico evaluation of primer specificity and secondary structure using Primer-BLAST and OligoAnalyzer. Nine primer pairs were initially generated and evaluated based on primer length, %GC, Tm, self3’ complementarity and amplicon size. Secondary structure analysis revealed strong self-dimer formation in pair 8, whereas pair 2 showed weak secondary structure within acceptable ΔG threshold (-9 kcal/mol). Experimental validation was conducted throught gradient PCR to optimize annealing temperature, followed by agarose gel 2%. Primer 2_CaPIP produced specific and clear amplification, with an optimal annealing temperature of 57.3oC. This study provides a validated CaPIP primer set suitable for qPCR-based gene expression analysis in C. annuum, supporting future molecular studies on drought stress tolerance.
Downloads
References
Afzal, Z., Howton, T., Sun, Y., & Mukhtar, M. (2016). The Roles of Aquaporins in Plant Stress Responses. Journal of Developmental Biology, 4(1), 9. https://doi.org/10.3390/jdb4010009
AlHarbi, A., Saleh, A., Al-Omran, A., & Wahb-Allah, M. (2014). Response of Bell-Pepper (Capsicum annuum L.) to Salt Stress and Deficit Irrigation Strategy Under Greenhouse Conditions. Acta Horticulturae, 1034, 443–450. https://doi.org/10.17660/actahortic.2014.1034.54
Aurora, D. A., Novarino, W., Tjong, D. H., Dahelmi, D., Syaifullah, S., Setiawan, A., & Roesma, D. I. (2025). Primer Design of Sumatran Striped Rabbit (Nesolagus netscheri Schlegel, 1880) using Primer-BLAST and AliView Program. Jurnal Biologi Tropis, 25(1), 599–605. https://doi.org/10.29303/jbt.v25i1.8499
Baker, P., Hill, P. J., Snarr, B. D., Alnabelseya, N., Pestrak, M. J., Lee, M. J., Jennings, L. K., Tam, J., Melnyk, R. A., Parsek, M. R., Sheppard, D. C., Wozniak, D. J., & Howell, P. L. (2016). Exopolysaccharide biosynthetic glycoside hydrolases can be utilized to disrupt and prevent Pseudomonas aeruginosa biofilms. Science Advances, 2(5), e1501632. https://doi.org/10.1126/sciadv.1501632
Bustin, S., & Huggett, J. (2017). qPCR primer design revisited. Biomolecular Detection and Quantification, 14, 19–28. https://doi.org/10.1016/j.bdq.2017.11.001
Bustin, S. A., & Mueller, R. (2005). Real-time reverse transcription PCR (qRT-PCR) and its potential use in clinical diagnosis. Clinical Science, 109(4), 365–379. https://doi.org/10.1042/cs20050086
Chen, Z., Halford, N. G., & Liu, C. (2023). Real-Time Quantitative PCR: Primer Design, Reference Gene Selection, Calculations and Statistics. Metabolites, 13(7), 806. https://doi.org/10.3390/metabo13070806
Chuang, L., Cheng, Y., & Yang, C. (2013). Specific primer design for the polymerase chain reaction. Biotechnology Letters, 35(10), 1541–1549. https://doi.org/10.1007/s10529-013-1249-8
Deshmukh, R. K., Sonah, H., & Bélanger, R. R. (2016). Plant Aquaporins: Genome-Wide Identification, Transcriptomics, Proteomics, and Advanced Analytical Tools. Frontiers in Plant Science, 7, 1896. https://doi.org/10.3389/fpls.2016.01896
Fakih, T. M., Wijaya, S., & Priani, S. E. (2021). Desain Primer Gen 12S sRNA dari DNA Mitrokondria Babi (Sus scrofa) secara In Silico sebagai Kandidat Primer dalam Analisis Molekuler Kehalalan Produk. Jurnal Sains Farmasi & Klinis, 8(3), 316. https://doi.org/10.25077/jsfk.8.3.316-322.2021
Fraige, K., Travensolo, R. F., & Carrilho, E. (2013). Analysis of seven STR human loci for paternity testing by microchip electrophoresis. Brazilian Archives of Biology and Technology, 56(2), 213–221. https://doi.org/10.1590/s1516-89132013000200006
Handoyo, D., & Rudiretna, A. (2000). General Principles and Implementation of Polymerase Chain Reaction. Unitas, 9(1). http://repository.ubaya.ac.id/35/1/ART002.pdf
Li, Z., Chen, J., Hao, Y., & Nair, S. K. (2012). Structures of the PelD Cyclic Diguanylate Effector Involved in Pellicle Formation in Pseudomonas aeruginosa PAO1. Journal of Biological Chemistry, 287(36), 30191–30204. https://doi.org/10.1074/jbc.m112.378273
Masnaini, M., Achyar, A., Chatri, M., Putri, D. H., Ahda, Y., & Irdawati. (2023). Primer Design and Optimization of PCR Methods for Detecting Mixed Rat Meat in Food Samples. In Advances in biological sciences research/Advances in Biological Sciences Research (pp. 282–289). https://doi.org/10.2991/978-94-6463-166-1_37
Meagher, R. J., Priye, A., Light, Y. K., Huang, C., & Wang, E. (2018). Impact of primer dimers and self-amplifying hairpins on reverse transcription loop-mediated isothermal amplification detection of viral RNA. The Analyst, 143(8), 1924–1933. https://doi.org/10.1039/c7an01897e
Mubarak, S. M. H., Al-Koofee, D. A. F., Radhi, O. A., Ismael, J. M., & Al-Zubaidi, Z. F. (2020). An optimization and common troubleshooting solving in polymerase chain reaction technique. Systematic Reviews in Pharmacy, 11(2), 427–436. https://doi.org/10.5530/srp.2020.2.63
Ntanasi, T., Karavidas, I., Savvas, D., Spyrou, G. P., Giannothanasis, E., Consentino, B. B., Papasotiropoulos, V., Sabatino, L., & Ntatsi, G. (2025). Physiological and Yield Responses of Pepper (Capsicum annuum L.) Genotypes to Drought Stress. Plants, 14(13), 1934. https://doi.org/10.3390/plants14131934
Oktavianti, R. (2019). The Application of PCR (Polymerase Chain Reaction) Using Specific Primer to Detect Chillies Drought Tolerant. Jurnal Agronomi Tanaman Tropika, 1(2), 49–66. https://doi.org/10.36378/juatika.v1i2.176
Pradnyaniti, D. G., Wirajana, I. N., & Yowani, S. C. (2013). Desain primer secara in silico untuk amplifikasi fragmen gen RPOB Mycobacterium tuberculosis dengan polymerase chain reaction (PCR). Jurnal Farmasi Udayana, 2(3), 279788. https://www.neliti.com/publications/279788/desain-primer-secara-in-silico-untuk-amplifikasi-fragmen-gen-rpob-mycobacterium
SantaLucia, J. (2007). Physical principles and Visual-OMP software for optimal PCR design. Methods in Molecular Biology, 402, 3–33. https://doi.org/10.1007/978-1-59745-528-2_1
Sari, E. N. (2018). Desain Primer Untuk Amplifikasi Regio Promoter Gen Inha Isolat P016 Multidrug Resistance Mycobacterium Tuberculosis Dengan Metode Polymerase Chain Reaction. Jurnal Farmasi Udayana, 34. https://doi.org/10.24843/jfu.2018.v07.i01.p01
Sasmito, D. E. K., Kurniawan, R., & Muhimmah, I. (2014). Karakteristik Primer pada Polymerase Chain Reaction (PCR) untuk Sekuensing DNA: Mini Review. Seminar Nasional Informatika Medis (SNIMed). https://journal.uii.ac.id/snimed/article/view/6329
Syamsidi, A., Aanisah, N., Fiqram, R., & Jultri, I. A. (2021). Primer Design and Analysis for Detection of mecA gene. Journal of Tropical Pharmacy and Chemistry, 5(3), 245–253. https://doi.org/10.25026/jtpc.v5i3.297
Syamsurizal, S., Handayani, D., Kadri, H., & Badriyya, E. (2019). Genotyping SNP rs7903146 TCF7L2 gene for detection T2DM in Indonesian melayu ethnic. Journal of Physics Conference Series, 1317(1), 012090. https://doi.org/10.1088/1742-6596/1317/1/012090
Wang C. (2016). Primer Design. Centre for Medical Parasitology University of Copenhagen. Health Science, Denmark
Yang, Z., Le, J. T., Hutter, D., Bradley, K. M., Overton, B. R., McLendon, C., & Benner, S. A. (2020). Eliminating primer dimers and improving SNP detection using self-avoiding molecular recognition systems. Biology Methods and Protocols, 5(1), bpaa004. https://doi.org/10.1093/biomethods/bpaa004
Ye, J., Coulouris, G., Zaretskaya, I., Cutcutache, I., Rozen, S., & Madden, T. L. (2012). Primer-BLAST: A tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics, 13(1), 134. https://doi.org/10.1186/1471-2105-13-134
Zhang, S., Feng, M., Chen, W., Zhou, X., Lu, J., Wang, Y., Li, Y., Jiang, C., Gan, S., Ma, N., & Gao, J. (2019). In rose, transcription factor PTM balances growth and drought survival via PIP2;1 aquaporin. Nature Plants, 5(3), 290–299. https://doi.org/10.1038/s41477-019-0376-1
Zhang, G., Zhang, Z., Wan, Q., Zhou, H., Jiao, M., Zheng, H., Lu, Y., Rao, S., Wu, G., Chen, J., Yan, F., Peng, J., & Wu, J. (2023). Selection and Validation of Reference Genes for RT-qPCR Analysis of Gene Expression in Nicotiana benthamiana upon Single Infections by 11 Positive-Sense Single-Stranded RNA Viruses from Four Genera. Plants, 12(4), 857. https://doi.org/10.3390/plants12040857.
Submitted
Accepted
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Bioeksperimen: Jurnal Penelitian Biologi

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.











